About me
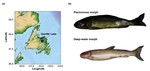
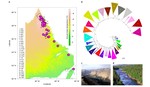
Hello I’m Cam Nugent, a Research Scientist at Fisheries and Oceans Canada. I am a bioinformatician whose work integrates aspects of data science, machine learning, and software development in the study of evolutionary genomics. My current research aims to characterize the structure of wild populations of ecologically and economically important aquatic species found in Atlantic Canada. I use large-scale genomic data, environmental information, statistics, and machine learning methodologies to understand the structure of wild populations and characterize their susceptibility to the effects of climate change. These discoveries are helping Fisheries and Oceans Canada to manage and protect Canada’s aquatic ecosystems and fisheries.
Interests
- Data science
- Machine learning
- Software development
- Statistics
- Bioinformatics
- Genomics
- Evolutionary biology
- Biological applications of machine learning algorithms
Education
-
Ph.D., 2019
University of Guelph, Department of Integrative Biology
-
B.Sc. (Hons), 2014
Queen's University